Blinda's Class

Classwork for BIMM143
Lab 07 Machine Learning 1
Blinda Sui (PID: A17117043)
Today we will explore some fundamental machine learning methods including clustering and dimensionality reduction.
K-means clustering
To see how this works let’s first makeup some data to cluster where we
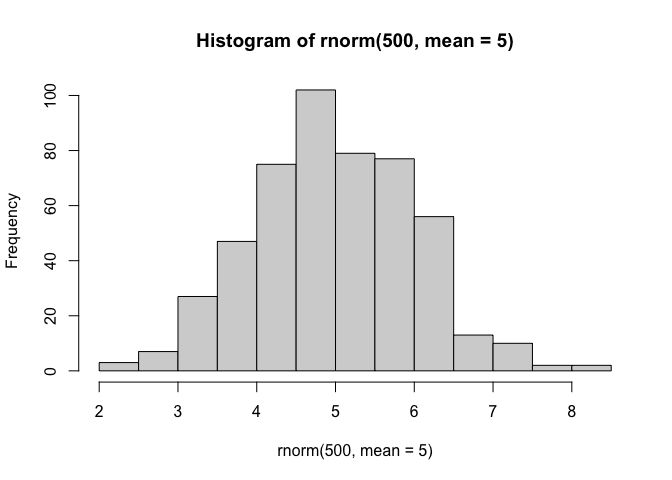
know what the answer should be. We can use the rnorm() function to
help here:
hist(rnorm(500, mean=5))
x <- c(rnorm(30, mean=-3),rnorm(30, mean=3))
y <- rev(x) #reverse x
cbind(x, y)
x y
[1,] -3.558170 3.632061
[2,] -1.501385 3.710897
[3,] -4.074267 4.027479
[4,] -3.679126 4.299014
[5,] -3.804401 4.355379
[6,] -3.319883 3.243559
[7,] -3.832704 3.013477
[8,] -4.289685 3.463544
[9,] -1.637985 1.816822
[10,] -4.374156 2.901404
[11,] -3.204849 3.673818
[12,] -4.145656 2.803598
[13,] -1.541535 2.742934
[14,] -3.476352 2.463249
[15,] -4.208345 3.008345
[16,] -3.346022 2.490190
[17,] -3.906891 1.085147
[18,] -4.125254 2.995049
[19,] -2.027630 5.267389
[20,] -2.417331 3.188682
[21,] -1.634382 2.361781
[22,] -2.590232 1.942936
[23,] -2.093131 3.225211
[24,] -5.301714 2.437554
[25,] -3.442146 3.071399
[26,] -2.689639 3.458139
[27,] -4.147775 1.796105
[28,] -4.277262 3.103786
[29,] -4.229445 3.100339
[30,] -2.141675 1.330859
[31,] 1.330859 -2.141675
[32,] 3.100339 -4.229445
[33,] 3.103786 -4.277262
[34,] 1.796105 -4.147775
[35,] 3.458139 -2.689639
[36,] 3.071399 -3.442146
[37,] 2.437554 -5.301714
[38,] 3.225211 -2.093131
[39,] 1.942936 -2.590232
[40,] 2.361781 -1.634382
[41,] 3.188682 -2.417331
[42,] 5.267389 -2.027630
[43,] 2.995049 -4.125254
[44,] 1.085147 -3.906891
[45,] 2.490190 -3.346022
[46,] 3.008345 -4.208345
[47,] 2.463249 -3.476352
[48,] 2.742934 -1.541535
[49,] 2.803598 -4.145656
[50,] 3.673818 -3.204849
[51,] 2.901404 -4.374156
[52,] 1.816822 -1.637985
[53,] 3.463544 -4.289685
[54,] 3.013477 -3.832704
[55,] 3.243559 -3.319883
[56,] 4.355379 -3.804401
[57,] 4.299014 -3.679126
[58,] 4.027479 -4.074267
[59,] 3.710897 -1.501385
[60,] 3.632061 -3.558170
plot(x)
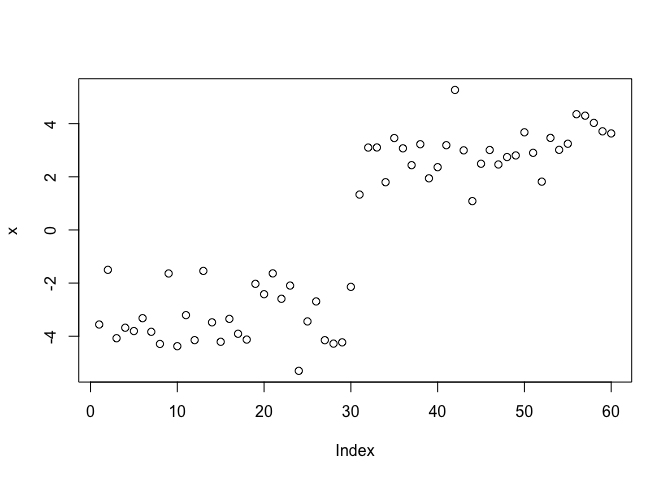
The function for K-means clustering in “base” R is kmeans()
kmeans(x, centers = 2)
K-means clustering with 2 clusters of sizes 30, 30
Cluster means:
[,1]
1 3.000338
2 -3.300634
Clustering vector:
[1] 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 1 1 1 1 1 1 1 1
[39] 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
Within cluster sum of squares by cluster:
[1] 23.48218 30.61468
(between_SS / total_SS = 91.7 %)
Available components:
[1] "cluster" "centers" "totss" "withinss" "tot.withinss"
[6] "betweenss" "size" "iter" "ifault"
k <- kmeans(x, centers = 2)
k
K-means clustering with 2 clusters of sizes 30, 30
Cluster means:
[,1]
1 3.000338
2 -3.300634
Clustering vector:
[1] 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 1 1 1 1 1 1 1 1
[39] 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
Within cluster sum of squares by cluster:
[1] 23.48218 30.61468
(between_SS / total_SS = 91.7 %)
Available components:
[1] "cluster" "centers" "totss" "withinss" "tot.withinss"
[6] "betweenss" "size" "iter" "ifault"
#:) Clustering vector: the first 30 belongs to cluster 1 and last 30 belongs to cluster 2 #:) Within cluster sume of squares by cluster (WCSS): For k=2 clusters, these are the WCSS values for cluster 1 and cluster 2. Lower = tighter cluster. Here, cluster 2 is looser (30.12 > 16.66). #:) (between_SS / total_SS = 92.4%): the proportion of total variance “explained” by the separation between the two cluster centroids = only 7.6% of the variance remains within clusters.
To get the results of the returned list object we can use the dollar $
syntax.
Q. How many points are in each cluster?
k$size
[1] 30 30
Q. What ‘component’ of your result object details - cluster assignment/membership? - cluster center?
k$cluster
[1] 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 1 1 1 1 1 1 1 1
[39] 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
k$centers
[,1]
1 3.000338
2 -3.300634
Q. Make a clustering results figure of the data colored by cluster membership and show cluster centers.
plot(x, col = c("red", "blue"))
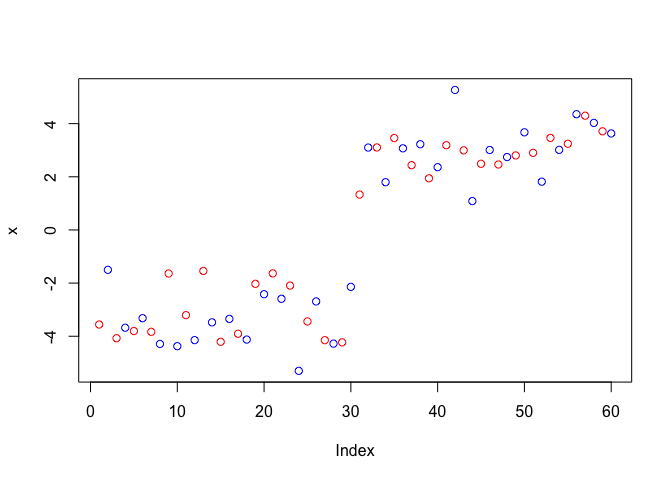
plot(x, col = 7)
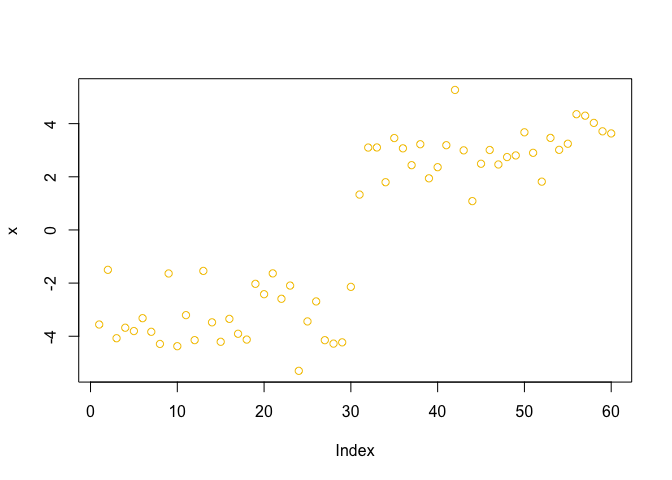
plot(x, col = k$cluster, pch=16)
points(k$centers, col="blue", pch=15, cex=2)
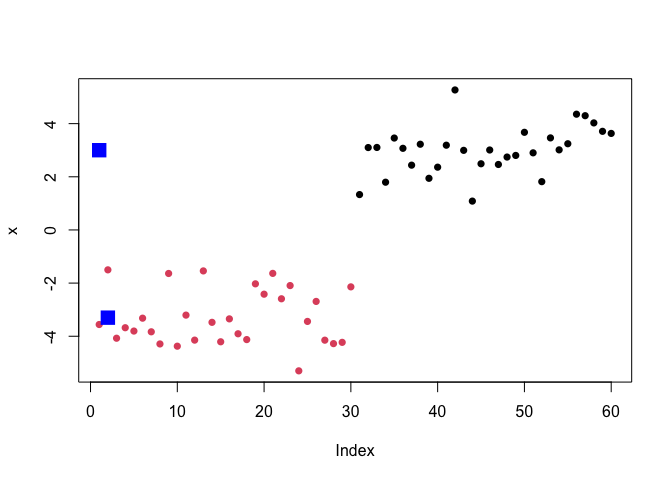
K-means clustering is very popular as it is very fast and relatively straight forward: it takes numeric data as input and returns the clusterm membership vector etc.
The “issue” is we tell kmeans() how many clusters we want!
Q. Run kmeans again and cluster into 4 frps/clusters and plot the results like we did above?
k4 <- kmeans(x, centers = 4)
plot(x, col=k4$cluster)
points(k4$centers, pch=15)
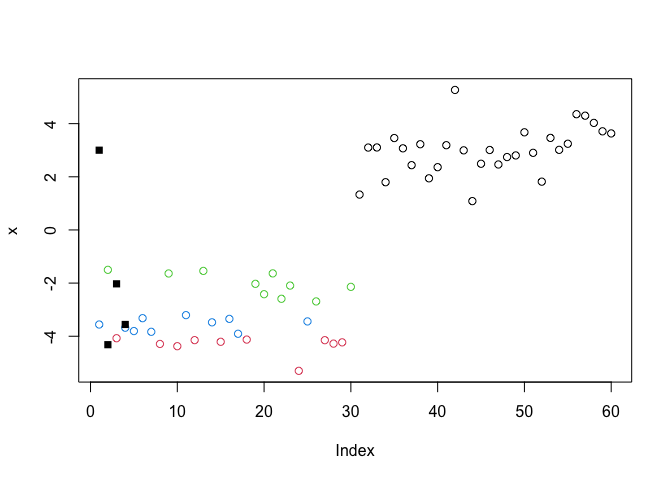
Scree plot to pick k center value
brute-force
k1 <- kmeans(x, centers = 1)
k2 <- kmeans(x, centers = 2)
k3 <- kmeans(x, centers = 3)
k4 <- kmeans(x, centers = 4)
k5 <- kmeans(x, centers = 5)
z <- c(k1$tot.withinss,
k2$tot.withinss,
k3$tot.withinss,
k4$tot.withinss,
k5$tot.withinss)
plot(z, typ = "b")
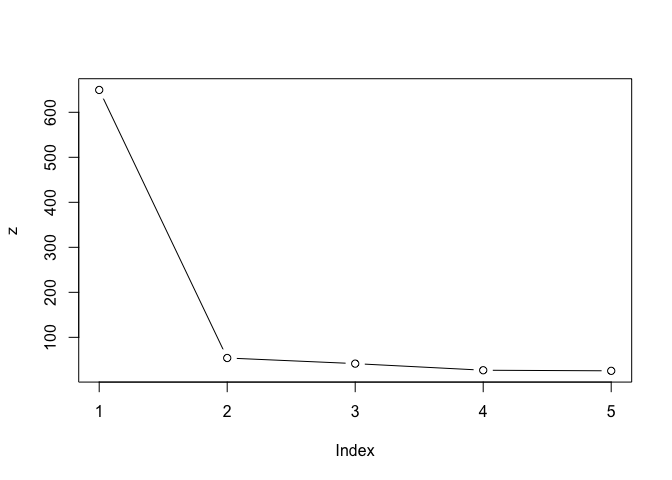
n <- NULL
for(i in 1:5) {
n <- c(n, kmeans(x, centers = i)$tot.withinss)
}
plot(n, typ = "b")
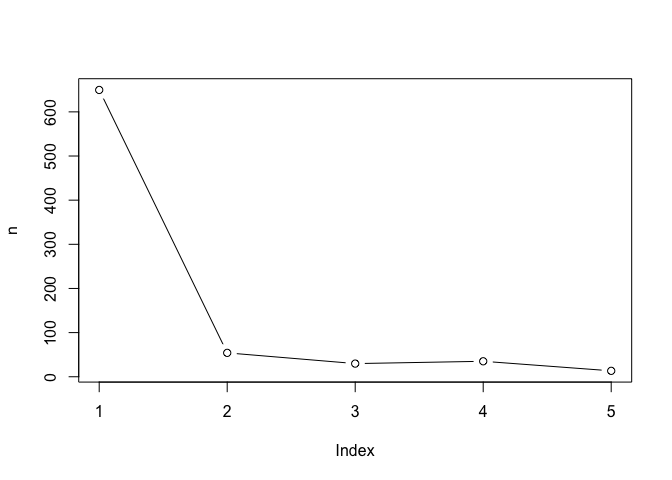
Hierarchical CLustering
The main “base” R function for Hierarchical Clustering is called
cluster(). Here we can’t just input our data, we need to first
calculate a distance matrix (e.g.dist()) for our data and use this as
input to hclust()
d <- dist(x)
hc <- hclust(d)
hc
Call:
hclust(d = d)
Cluster method : complete
Distance : euclidean
Number of objects: 60
There is a plot method for hclust results, lets try it
plot(hc)
abline(h=8, col="red") #abline(): to add straight lines to an existing plot.
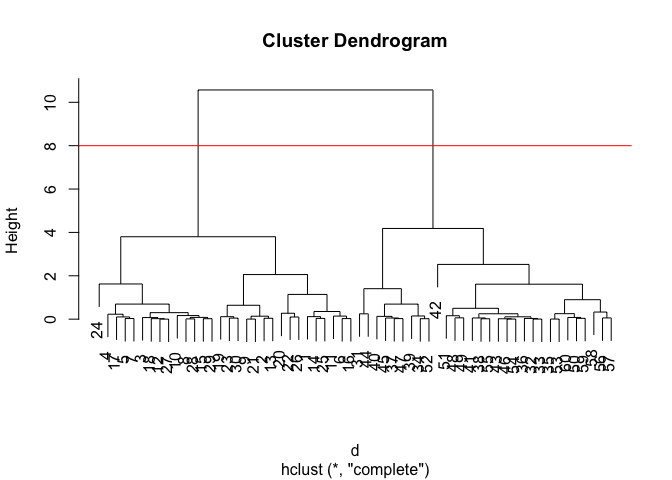
To get our cluster “membership” vector (i.e. our main clustering result) we can “cut” the tree at a given height or at height that yields a given “k” gorups.
cutree(hc, h=8)
[1] 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2
[39] 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2
grps <- cutree(hc, k=2)
Q. Plot the data with our hclust result coloring
plot(x, col=grps)
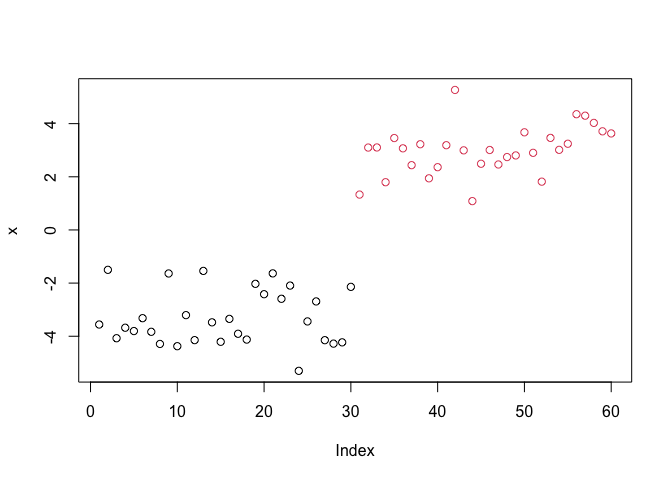
Principal Component Analysis (PCA)
PCA of UK food data
Import food data from an online CSV file:
url <- "https://tinyurl.com/UK-foods"
x <- read.csv(url)
head(x)
X England Wales Scotland N.Ireland
1 Cheese 105 103 103 66
2 Carcass_meat 245 227 242 267
3 Other_meat 685 803 750 586
4 Fish 147 160 122 93
5 Fats_and_oils 193 235 184 209
6 Sugars 156 175 147 139
rownames(x) <- x[, 1]
x <- x[, -1]
x
England Wales Scotland N.Ireland
Cheese 105 103 103 66
Carcass_meat 245 227 242 267
Other_meat 685 803 750 586
Fish 147 160 122 93
Fats_and_oils 193 235 184 209
Sugars 156 175 147 139
Fresh_potatoes 720 874 566 1033
Fresh_Veg 253 265 171 143
Other_Veg 488 570 418 355
Processed_potatoes 198 203 220 187
Processed_Veg 360 365 337 334
Fresh_fruit 1102 1137 957 674
Cereals 1472 1582 1462 1494
Beverages 57 73 53 47
Soft_drinks 1374 1256 1572 1506
Alcoholic_drinks 375 475 458 135
Confectionery 54 64 62 41
x <- read.csv(url, row.names = 1)
x
England Wales Scotland N.Ireland
Cheese 105 103 103 66
Carcass_meat 245 227 242 267
Other_meat 685 803 750 586
Fish 147 160 122 93
Fats_and_oils 193 235 184 209
Sugars 156 175 147 139
Fresh_potatoes 720 874 566 1033
Fresh_Veg 253 265 171 143
Other_Veg 488 570 418 355
Processed_potatoes 198 203 220 187
Processed_Veg 360 365 337 334
Fresh_fruit 1102 1137 957 674
Cereals 1472 1582 1462 1494
Beverages 57 73 53 47
Soft_drinks 1374 1256 1572 1506
Alcoholic_drinks 375 475 458 135
Confectionery 54 64 62 41
some base figures
barplot(as.matrix(x), beside=T, col=rainbow(nrow(x)))
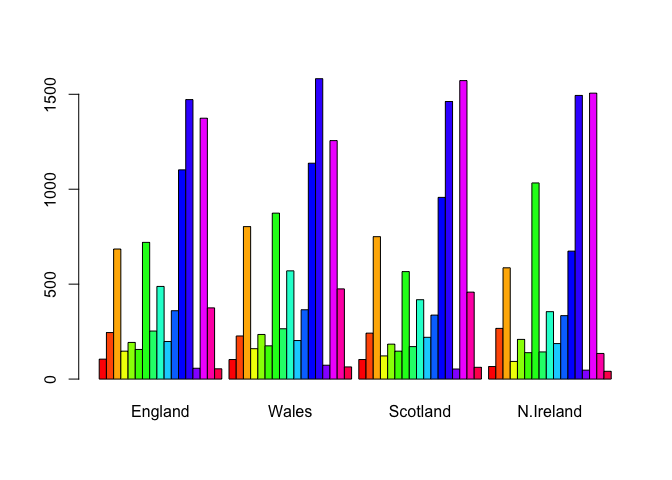
barplot(as.matrix(x), beside=F, col=rainbow(nrow(x)))
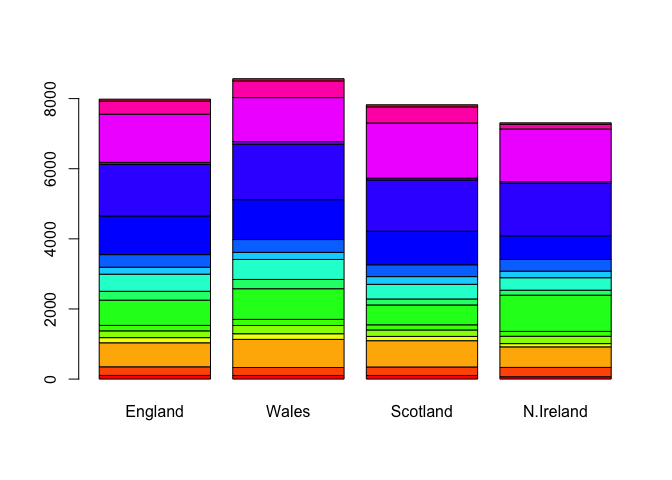
There is one plot that can be useful for small datasets:
pairs(x, col=rainbow(nrow(x)), pch=16)
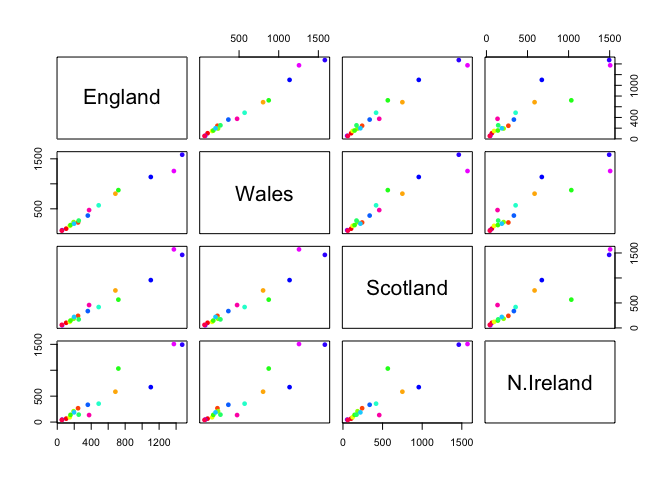
Main point: It can be difficult to spot major trends and patterns even in relatively small multivariate datasets (here we only have 17 dimensions, typically we have 1000s).
PCA to the rescue
The main function in “base” R for PCA is called prcomp()
I will take the transpose of our data so the “food” are in the columns:
pca <- prcomp(t(x))
summary(pca)
Importance of components:
PC1 PC2 PC3 PC4
Standard deviation 324.1502 212.7478 73.87622 2.921e-14
Proportion of Variance 0.6744 0.2905 0.03503 0.000e+00
Cumulative Proportion 0.6744 0.9650 1.00000 1.000e+00
cols <- c("orange", "red", "blue", "darkgreen")
plot(pca$x[,1], pca$x[,2], col=cols, pch=16)
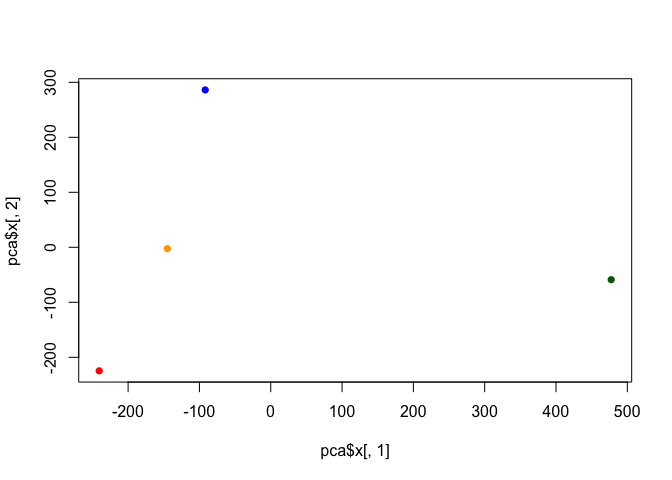
library(ggplot2)
ggplot(pca$x) +
aes(PC1, PC2) +
geom_point(col=cols)
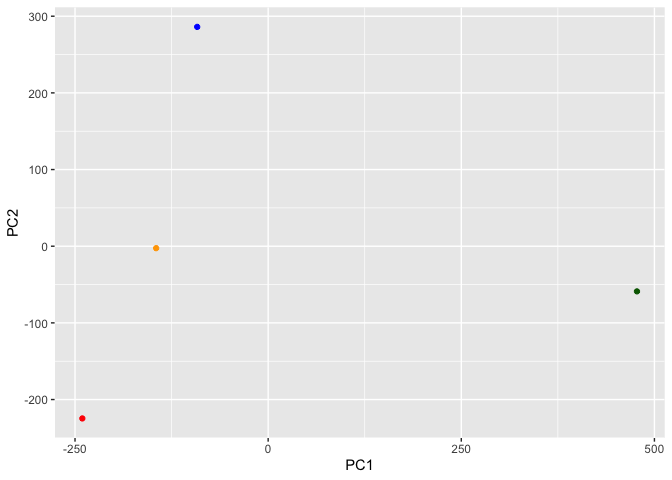
ggplot(pca$rotation) +
aes(PC1, rownames(pca$rotation)) +
geom_col()
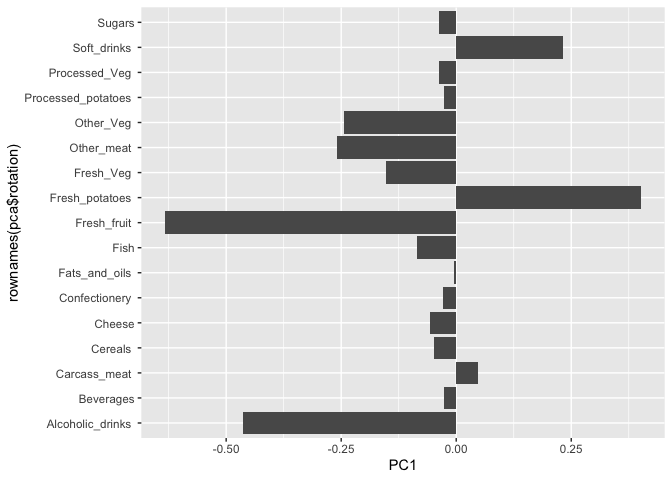
PCA looks super useful and we will come back to describe this further next day ;-)